Archive for category Journals Publishing
DITA – A framework for scientific publishing?
Posted by peanutbutter in data standards, Journals Publishing on February 1, 2011
There are two industry recognised standards for XML based documentation. These are Docbook and DITA (Darwin Information Typing Architecture).
Docbook is the older of the two specifications and created specifically for technical documentation. DITA, is a younger specification which grew out of IBM, and is referred to as having its own architecture and was designed to provide structure to more than just a book. Both specifications are OASIS standards.
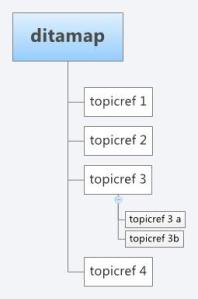
As with XML schemas, both specifications can be extended to include bespoke features. However, Docbook is based more on a book structure with Sections and subsections, where as DITA is built around topics that can be built up in any arrangement based on a document map. A DITA topic is open to specialisation itself, however, a topic has only three required elements
- An id attribute
- A title
- A body
- ,
A topic can exist as a single XML file which can be composed into any arrangement for publication through the use of a document map. A DITA structure would present a more flexible architecture where the same “topic”, i.e a journal article section, such as an abstract, materials and methods, or results, could be included with ease more than one publication, correctly referenced. In this respect DITA is more like an object-oriented document schema, and can be more easily repurposed (in terms of structure) for any output format (i.e pdf, HTML). In the same respect, Docbook can be configured with some work to behave on a more topic by topic basis and DITA can support a book based methodology. They are after all both XML schemas and are equally extensible or open to specialisation.
As its a standard, whole ecosystems have emerged which makes use of the DITA architecture. For example, DITA for publishers provides libraries to convert DITA markup into HTML, PDF, EPUB, and Kindle rendering support. This allows content structures in DITA to be repurposed for different audiences or different devices with relative ease.
I have recently started using DITA as an architecture to represent content, primarily designed for books. However, with new demands appearing for different delivery mechanisms of the traditional textbook, such as Web delivery and ebooks, DITA is proving to be immensely powerful to deliver the same content through different mediums with relative ease and speed. In using it, it seems obvious that a DITA architecture would benefit the representation of content within a journal article, allowing references re-purposing and multiple format delivery. Maybe a topic for discussion through the Beyond the PDF forum.
In the end, it’s just XML, so I wont repeat the virtues of content markup through XML. However, for me its main advantage is the object oriented -like topic structure as a working architecture.
Related articles
- Using RDFa with DITA and DocBook (devx.com)
- Dita Educational Use cases (docs.oasis-open.org)
- Converting documents between a wiki and Word, XML, FrameMaker or other help formats (ffeathers.wordpress.com)
- Future-proofing e-books with XML (teleread.com)
- The PDF Landscape for DITA Content (tc.eserver.org)
Is a knol a scientific publishing platform?
Posted by peanutbutter in bioinformatics, Journals Publishing, open data, open science, Social Media on August 19, 2008
 Image via CrunchBase, source unknown
Image via CrunchBase, source unknown Google has recently released knol, which most people are calling Google’s version of Wikipedia. The main difference between a knol and a wikipedia article is that a knol has associated authorship or ownership against an article. This factor has caused some issues an outcries focusing on the merits of the wisdom of crowds verses the merits of single individuals and the whole ethos if information dissemination on the Web. (There are too many to cite but some discussion can be found on FriendFeed+knol)
However, on looking at knol and having a snoop around I was not drawn into thinking about competing with Wikipedia or advertising revenue, rather what struck me is that a knol, with owner authorship looks incredibly like a scientific journal publication platform…
According to the Introduction to knol (or instructions for authors) you to can write about anything you like, so disseminiation of science must fall under that. You can collaborate on a unit of knowledge (or manuscript) with other authors and they are listed – I assume the contribution to the text will also be stored in the revision history, makes the authors contribution section a little easier to write. It is not limited to one article per subject, so this allows all manner of opinions or scientific findings to be reported contrasted and compared with each other. You can select your copyright and license for your article (rather than handing it over). You can request a (peer-) review of the article, However more important the article is available for continuous peer-review in the form of comments on the article.
So is a knol a Google Wikipedia or is it a scientific publishing platform? What would prevent publishiing a knol and getting credit (hyperlinks), citations (analytics) impact factors (page ranks) in the same way you do for the traditional scientific publishers? You would of course not have to pay for the privilage of trying to diseminate your findings, loosing copyright and then asking your institution to pay for a subscription so you and your lab members can read your own articles. In fact you, as an author (lab, institution) can even share revenue for your article via adsense.
Some traditional publishers are trying to embrace new mechanisms of disseminating scientific knowledge. Only today Nature Reviews Genetics (as described by Nacsent) published the paper Towards a cyberinfrastructure for the biological sciences: progress, visions and challenges by Lincoln Stein and published the supplementary material as a community editable wiki.
With knol as a scientific publishing platform, what can the traditional scientific publishing houses now offer for the publishing fees? Faster turn around? revenue sharing? Are they really still the gatekeepers of scientific knowledge? Or in the Web 2.0 era has that mantle passed to Google? Certainly, in the first instance it would make a nice preceedings platform.
MIAPE: Gel Informatics is now available for Public Comment
Posted by peanutbutter in bioinformatics, data standards, Journals Publishing, MIBBI, Proteomics on August 18, 2008

PSI logo
The MIAPE: Gel Informatics module formalised by the Proteomics Standards Initiative (PSI) now available for Public Comment on the PSI Web site. Typically alot of this information will be contained in the image analysis software, so we would especially encourage software vendors to review the document. The public
comment period enables the wider community to provide feedback on a proposed standard before it is formally accepted, and thus is an important step in the standardisation process.
This message is to encourage you to contribute to the standards development activity by commenting on the material that is available online. We invite both positive and negative comments. If negative comments are being made, these could be on the relevance, clarity, correctness, appropriateness, etc, of the proposal as a whole or of specific parts of the proposal.
If you do not feel well placed to comment on this document, but know someone who may be, please consider forwarding this request. There is no requirement that people commenting should have had any prior contact with the PSI.
If you have comments that you would like to make but would prefer not to make public, please email the PSI editor Norman Paton.
Double standards in Nature biotechnology
Posted by peanutbutter in bioinformatics, data standards, Journals Publishing, MIBBI on August 7, 2008
OK, So that is a relatively inflammatory and controversial headline, edging on the side of tabloid sensationalism. What is refers to is probably a situation that I may never find myself in again, which is in this months edition of Nature Biotechnology I am an author on two, biological standards related publications.
The first is a letter advertising the PSI’s MIAPE Guidelines for reporting the use of gel electrophoresis in proteomics. This letter is also accompanied by letters referring to the MIAPE guidelines for Mass Spectrometry, Mass Spectrometry Informatics and protein modification data.
The second is a paper on the Minimum Information about a Biomedical or Biological Investigations (MIBBI) registry entitled Promoting coherent minimum reporting guidelines for biological and biomedical investigations: the MIBBI project.
The following press release describes this paper in more detail.
More than 20 grass-roots standardisation groups, led by scientists at the European Bioinformatics Institute (EMBL-EBI) and the Centre for Ecology & Hydrology (CEH), have combined forces to form the “Minimum Information about a Biomedical or Biological Investigation” (MIBBI) initiative. Their aim is to harmonise standards for high-throughput biology, and their methodology is described in a Commentary article, published today in the journal Nature Biotechnology.
Data standards are increasingly vital to scientific progress, as groups from around the world look to share their data and mine it more effectively. But the proliferation of projects to build “Minimum Information” checklists that describe experimental procedures was beginning to create problems. “There was no way of even finding all the current checklist projects without days of googling,” says the EMBL-EBI’s Chris Taylor, who shares first authorship of the paper with Dawn Field (CEH) and Susanna-Assunta Sansone (EMBL-EBI). “As a result, much of the great work that’s going into developing community standards was being overlooked, and different communities were at risk of developing mutually incompatible standards. MIBBI will help to prevent them from reinventing the wheel.
The MIBBI Portal already offers a one-stop shop for researchers, funders, journals and reviewers searching for a comprehensive list of minimum information checklists. The next step will be to build the MIBBI Foundry, which will bring together diverse communities to rationalise and streamline standardisation efforts. “Communities working together through MIBBI will produce non-overlapping minimal information modules,” says CEH’s Dawn Field. “The idea is that each checklist will fit neatly into a jigsaw, with each community being able to take the pieces that are relevant to them.” Some, such as checklists describing the nature of a biological sample used for an experiment, will be relevant to many communities, whereas others, such as standards for describing a flow cytometry experiment, may be developed and used by a subset of communities.
“MIBBI represents the first new effort taking the Open Biomedical Ontologies (OBO) as its role model”, says Susanna-Assunta Sansone. “The MIBBI Portal operates in a manner analogous to OBO as an open information resource, while the MIBBI Foundry fosters collaborative development and integration of checklists into self-contained modules just like the OBO Foundry does for the ontologies”.
There is a growing understanding of the value of such minimal information standards among biologists and an increased willingness to work together across disciplinary boundaries. The benefits include making experimental data more reproducible and allowing more powerful analyses over diverse sets of data. New checklist communities are encouraged to register with MIBBI and consider joining the MIBBI Foundry.
Press release issued by the EMBL-European Bioinformatics Institute and the Centre for Ecology and Hydrology, UK.
How do you select your Scientific Journal?
Posted by peanutbutter in bioinformatics, data standards, Journals Publishing, ontology, Proteomics on April 21, 2008
I was trying to work out a suitable journal to where I could submit a paper on sepCV, the PSI ontology for sample preparation and separation techniques. I found my self drawing up a table, so I thought I would blog it. My initial remit is that is should be in a proteomics relevant journal as well as bioinformatics, as we are trying to encourage a greater community contribution for term collection. In this respect it has to be open access. I would also prefer the journal to accept LateX instead on proprietary formats such as word. I was really disappointed with the number of journals that only accept word documents, even PlosOne refuses anything other than word or rft, tut, tut.
Based on this loose criteria, Proteome science comes top closely followed by Journal of Proteomics and Bioinformatics (if I sacrifice using word for open access). BMC Bioinformatics also ticks all the boxes but it misses out on the proteomics audience. The table below also includes Impact factor, but I did not really take that into consideration. Wouldn’t it be nice is there was an app that you could just enter in criteria like this, target audience, submission format, copyright, etc and get back the journals that meet these requirements. Something like this would save me an afternoon trawling through the web building spreadsheets.
How do you select your journal?
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Zotero library re-visioned
Posted by peanutbutter in backup, bioinformatics, howto, Journals Publishing, library, reference, Social Bookmarking, Social Media, subversion, zotero on February 22, 2008
I have been wanting to use Zotero now for a while for my reference library but could never work out how back up my library using subversion. My life is contained within subversion, I do not know how I could have possibly survived before all my work; code, presentations, papers, images and not to mention my thesis, is all perfectly backed up and re-visioned and floating happily in the cloud available to me from any machine. Zotero installs itself inside the firefox profile which makes it difficult to revision within the C:\\my-subversion” folder. What I decided to do was to create a new firefox profile (instructions here) within my-subversion folder then install zotero creating:
C:\\my-subversion\firefoxprofile\zotero
I then only added the zotero folder to my subversion repository. You could always revision your firefox profile but I decided not to. Now every time I add a new item to zotero the my-subversion folder indicates there has been a change and requires a commit. Obviously every time you add a pdf file to the library you will actually have to “SVN add” the file itself. This is not a problem for me as I try to keep my library light and not store to many pdfs.
I am also going to try and use zotero as an interface to my subversion repository, describing and tagging documents and code that I write, but more specifically presentations, so no more trying to work out what is contained in “Presentation1.ppt” or what file name I gave to that talk on data standards which I have to give tomorrow!
I am tagging my hard drive via Zotero, its just one big cloud.
goPubmed
Posted by peanutbutter in bioinformatics, Journals Publishing, ontology, search, semantic web, Social Media on January 6, 2008
In catching up with my reading lists of 2007 I was alerted to gopubmed via Deepak’s post. Gopubmed described itself as an ontology based literature search making use of both the Gene ontology and Mesh terms. There is also the ability to provide feedback or rather act as a curator for the search results. I have already noticed some mis-match in author details. In general though the interface is a vast improvement on pubmed’s tiered interface and the ability to refine the searches looks interesting. I have added gopubmed to my search engines within firefox and will have a play to see if it is any good. RSS feeds on search terms would be top of the wish list. Anybody used it in anger?
I see science
Posted by peanutbutter in bioinformatics, Journals Publishing, open data, open science, Social Media, Uncategorized, video on August 20, 2007
It is interesting to see new developments in the dissemination of scientific discourse, such as scientific blogging and paradigms such as open science, come on-line with developments in web-based social media. The latest medium to receive the Science 2.0 treatment (poor pun on applying Web 2.0 technologies for science) is the video.
YouTube is probably the granddaddy, or at least the most prominent of the video upload and broadcast services. Although YouTube doesn’t not have a defined science category, it is easy to find science related videos and lectures, mixed in with the general population. However, several specialist sites have appeared dealing specifically with science research, all have been labeled as “YouTube for science”. The most recent site is SciVee, which is a collaboration of the Public Library of Science (PLoS), the National Science Foundation (NSF) and the San Diego Supercomputer Center (SDSC). The fact the a publishing house (PLoS) has got involved in this effort is encouraging, and maybe an admittance that a paper, in isolation of public commentary, the data used to produce the paper, and a sensible presentation mechanism, is no longer sufficient in the web-based publishing era (or maybe I have got over excited and read to much into it). The most interesting feature on SciVee, and probably the most powerful compared to some of the other broadcasters, is the ability to link to an Open Access publication, setting the context and relevance of the video. The flip side could also be true, where a video provides evidence of the experiment, such as the methods or the displays the result.
Another scientific video broadcaster is the Journal of Visualized Experiments (JOVE). As the name suggests, JOVE focuses on capturing the experiment performed within the laboratory, rather than a presentation, or general scientific discourse. As a result JOVE can be though of as a visual protocol or methods journal and is stylaised as a traditional journal, already on Issue 6; a focus on Neuroscience.
If JOVE is a visual journal for life-science experiments then Bioscreencast could be thought of as a visual journal of Bioinformatics. Bioscreencast focusses on screencasts of software, providing a visual “How-to” on scientific software, presentations and demonstrations.
No doubt these three may not be the last scientific video publishers, but they have an opportunity to become well established ahead of the others. Now, where is my webcam, I need to video myself writing code and submit it to JOVE, produce a demo and submit it to Bioscreencast. Then I have to write a paper on it, submit the pre-prints to Nature Preccedings get it published in an Open Access Journal, then video myself giving a presentation on the paper, submit it to SciVee and link them all together.
Do scientists really believe in open science?
Posted by peanutbutter in bioinformatics, CARMEN, data standards, Journals Publishing, MIBBI, neuroinformatics, ontology, open data, open science, Social Media on June 26, 2007
I am writing this post as a collection of the current status and opinions of “Open Science”. The main reason being I have a new audience; I am working for the CARMEN e-Neuroscience project. This has exposed me, first hand, to a domain of the life-sciences to which data sharing and publicly exposing methodologies has not been readily adopted, largely it is claimed due to the size of the data in question and sensitive privacy issues.
Ascoli, 2006 also endorses this view of the neuroscience and offers some further reasons why this is the case . He also includes the example of exposing neuronal morphological data and argues the benefits and counters the reticence to sharing this type of data.
Hopefully, as the motivation for the CARMEN project is to store and share and facilitate the analysis of neuronal activity data, some of these issues can be overcome.
With this in mind I want to create this post to provide a collection of specific blogs, journal articles, relevant links and opinions which hopefully will be a jumping-off point to understanding the concept of Open Science and embracing the future methodologies in pushing the boundaries of scientific knowledge.
What is Open Science?
There is no hard and fast definition, although according to the Wikipedia entry:
“Open Science is a general term representing the application of various Open approaches (Open Source, Open Access, Open Data) to scientific endeavour. It can be partially represented by the Mertonian view of Science but more recently there are nuances of the Gift economy as in Open source culture applied to science. The term is in intermittent and somewhat variable use.”
“Open Science” encompasses the ideals of transparent working practices across all of the life-science domains, to share and further scientific knowledge. It can also be thought of to include the complete and persistent access to the original data from which knowledge and conclusions have been extracted. From the initial observations recorded in a lab-book to the peer-reviewed conclusions of a journal article.
The most comprehensive overview is presented by Bill Hooker over at 3quarks daily. He has written three sections under the title “The Future of Science is open”
In part 1, as the title suggests, Bill presents an overview on open access publishing and how this can lead to open-science (part 2). He suggests that
“For what I am calling Open Science to work, there are (I think) at least two further requirements: open standards, and open licensing.”
I don’t want to repeat the content already contained in these reviews, although I agree with Bill’s statement here. There is no point in having an open science philosophy if the data in question is not described or structured in a form that facilitates exchange, dissemination and evaluation of the data, hence the requirement of standards.
I am unaware of community endorsed standard reporting formats within Neuroscience. However, the proliferation of standards in Biology and Bioinformatics, is such, that it is fast becoming a niche domain in its own right. So much so, that there now exists a registry for Minimum Information reporting guidelines, following in the formats of MIAME and MIAPE. This registry is called MIBBI (Minimum Information for Biological and Biomedical Investigations) and aims to act as a “one-stop-shop” of existing standards life-science standards. MIBBI also provides a foundry where best practice for standards design can be fostered and disparate domains can integrate and agree on common representations of reporting guidelines for common technologies.
Complementary to standard data structures and minimum reporting requirements, is the terminology used to described the data; the metadata. Efforts are under way to standardise terminology which describes experiments, essential in an open environment, or simply in a collaboration. This is the goal of the Ontology of Biomedical Investigations (OBI) project which is developing “an integrated ontology for the description of biological and medical experiments and investigations. This includes a set of ‘universal’ terms, that are applicable across various biological and technological domains, and domain-specific terms relevant only to a given domain“. Already OBI is gaining momentum and currently supports diverse communities from Crop science to Neuroscience.
Open licensing of data may address the common arguments I hear for not releasing data, that “somebody might use it”, or the point blank refusal of “not until I publish my paper”. This is an unfortunate side effect of the “publish or perish” system as commented on bbgm and Seringhaus and Gerstein, 2007, and really comes down to due credit. In most cases this prevents real time assessment of research, complementary analysis or cross comparisons with other data sets to occur alongside the generation of the data, which would in no doubt enforce the validity of the research. Assigning computational amenable licenses to data, such as those proposed by Science Commons, maybe one way of ensuring that re-use of the data is always credited to the laboratory that generated the data. It is possible paradigm that “Data accreditation impact factors” could exist analogous to the impact factors of traditional peer-reviewed journals.
Open science may not just be be about releasing data associated with a peer-review journal, rather it starts from exposing the daily recordings and observations of an investigation, contained in the lab-book. One aspect of the “Open data” movement is that of “Open Notebook Science” a movement pioneered by Jean-Claude Bradley and the Useful Chemistry group, where their lab-book is is open and access-able on-line. This open notebook method was further discussed by a recent Nature editorial outlining the benefits of this approach. Exposing you lab-book could allow you to link the material and methods section of your publication, proving you actually did the work and facilitating the prospect of other researchers actually being able to repeat your ground breaking experiments.
Already many funders are considering data management or data sharing policies, to be applied to future research proposals. The BBSRC have recently released their data sharing policy which states that, “all research proposals submitted to BBSRC from 26th April 2007 must now include a statement on data sharing. This should include concise plans for data management and sharing or provide explicit reasons why data sharing is not possible or appropriate“. With these types of policies a requirement to research funding the “future of science is open“.
The “Open Science” philosophy appears to be gaining some momentum as is actively being discussed within the scientific blogosphere. This should not really come as a great surprise as science blogging can be seen as part of the “Open science” movement, openly sharing opinions and discourse. Some of the more prominent science blogs focusing on the open science ideal are Open access News, Michael Eisen’s Open Science Blog, Research Remix, Science Commons, Peter Murray-Rust.
There are of course alot more blogs discussing the issue. Performing an “open science search” on Postgenomic (rss feed on search terms please, Postgenomic) produces an up to the minute list of the open science discourse. Although early days, maybe even the “open science” group on Scintilla (still undecided on Scintilla) will be the place in the future for fostering the open science community.
According to Bowker’s description of the traditional model of scientific publishing, the journal article “forms the archive of scientific knowledge” and therefore there has been no need to hold on to the data after it has been “transformed” into a paper. This, incorporated with in-grained social fears, as a result of “publish or perish”, of not letting somebody see the experimental data before they get their peer-reviewed publication, will cripple the open science movement and slow down knowledge discovery. Computational amenable licences may go some way to solve this. But raising the awareness and a clear memorandum from the major journal publishers that, exposing real-time science and publishing data will not prevent publication as a peer-reviewed journal, can only help.
In synopsis I will quote Bill again as I think he presents a summary better than I could;
“My working hypothesis is that open, collaborative models should out-produce the current standard model of research, which involves a great deal of inefficiency in the form of secrecy and mistrust. Open science barely exists at the moment — infancy would be an overly optimistic term for its developmental state. Right now, one of the most important things open science advocates can do is find and support each other (and remember, openness is inclusive of a range of practices — there’s no purity test; we share a hypothesis not an ideology). “



![Reblog this post [with Zemanta]](https://i0.wp.com/img.zemanta.com/reblog_e.png)